VISION Research Projects – 1st cohort

A01 – Spatiotemporal orchestration of alphaherpesvirus capsid assembly, genome packaging and nuclear egress
Julian Kraft
Centre for Structural Systems Biology (CSSB)

A02 – Protein-protein interactions determining influenza A virus (IAV) shape and assembly
Konstantin Gilep
Centre for Structural Systems Biology (CSSB)
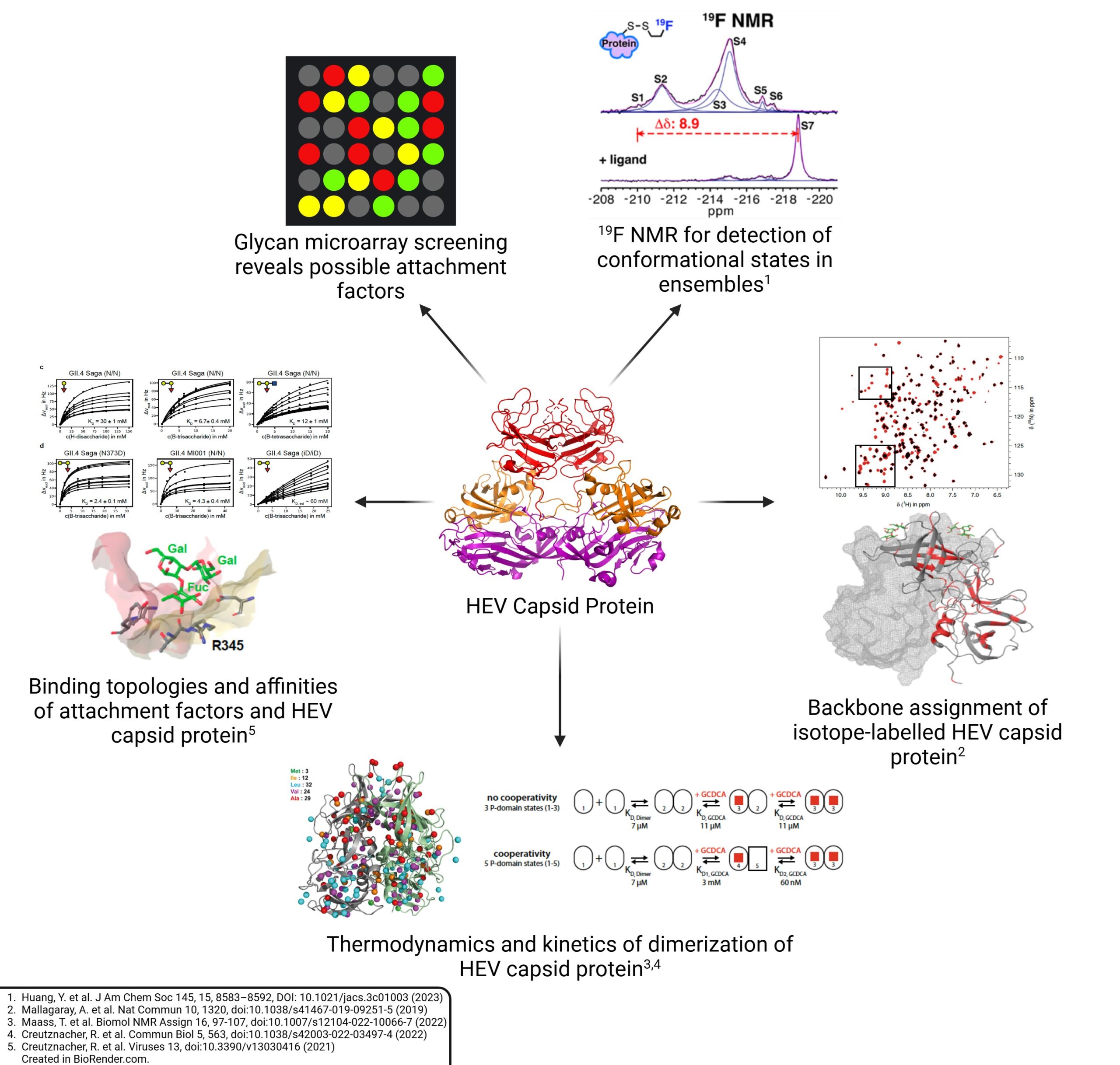
A03 – Probing interactions of Hepatitis E virus (HEV) capsid proteins with attachment factors and neutralizing antibodies using NMR
Nachiket Moti
Universität zu Lübeck (UzL)
–
–
–

A04 – From deciphering to blocking assembly of human norovirus-like particles
Jennifer Rothe
Centre for Structural Systems Biology (CSSB)
–
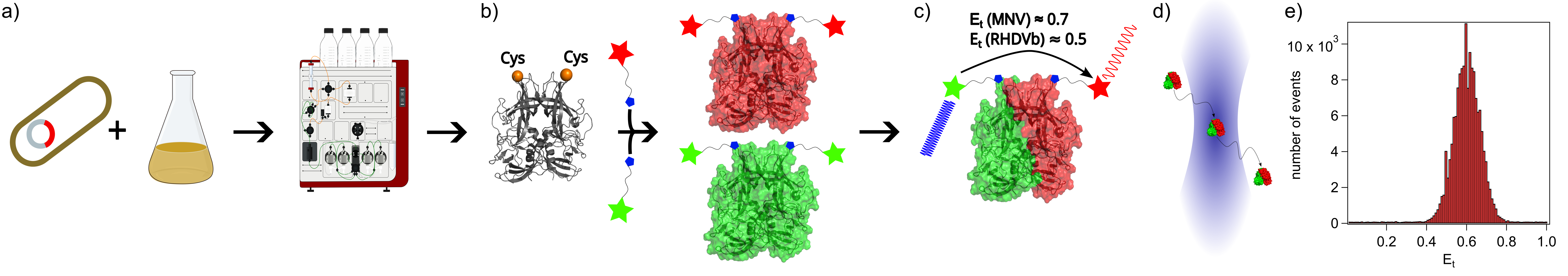
A05 – Dimerisation of viral capsid proteins investigated by single molecule FRET
Leon Westermann
associated
Universität zu Lübeck (UzL)

A06 – Secondary envelopment of herpesviruses
Jedrzej Mazur
associated
Centre for Structural Systems Biology (CSSB)
–
–
–
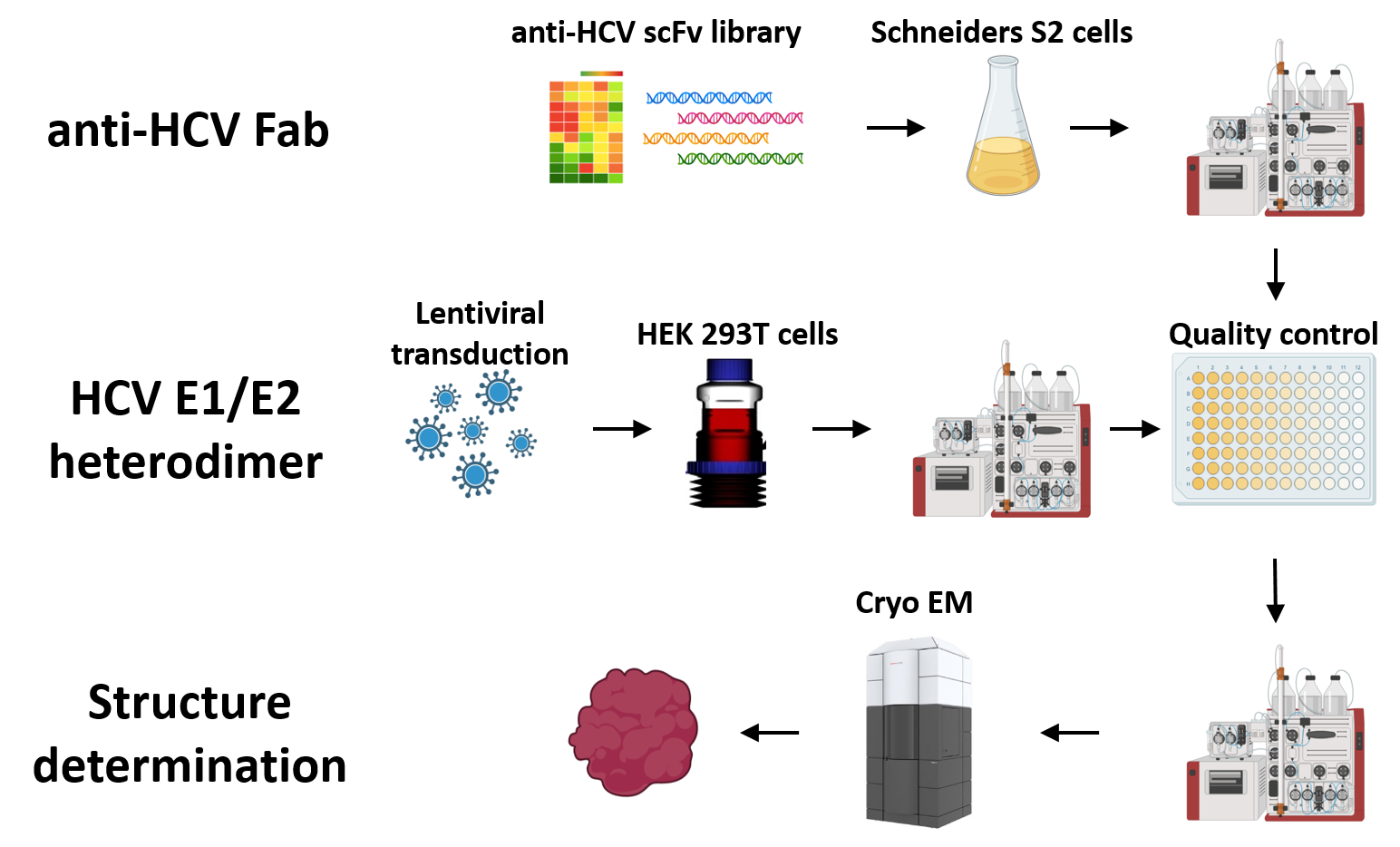
A07 – Structural Characterization of HCV-Antibody Interactions
Frithjof Besa
associated
Universität zu Lübeck (UzL)
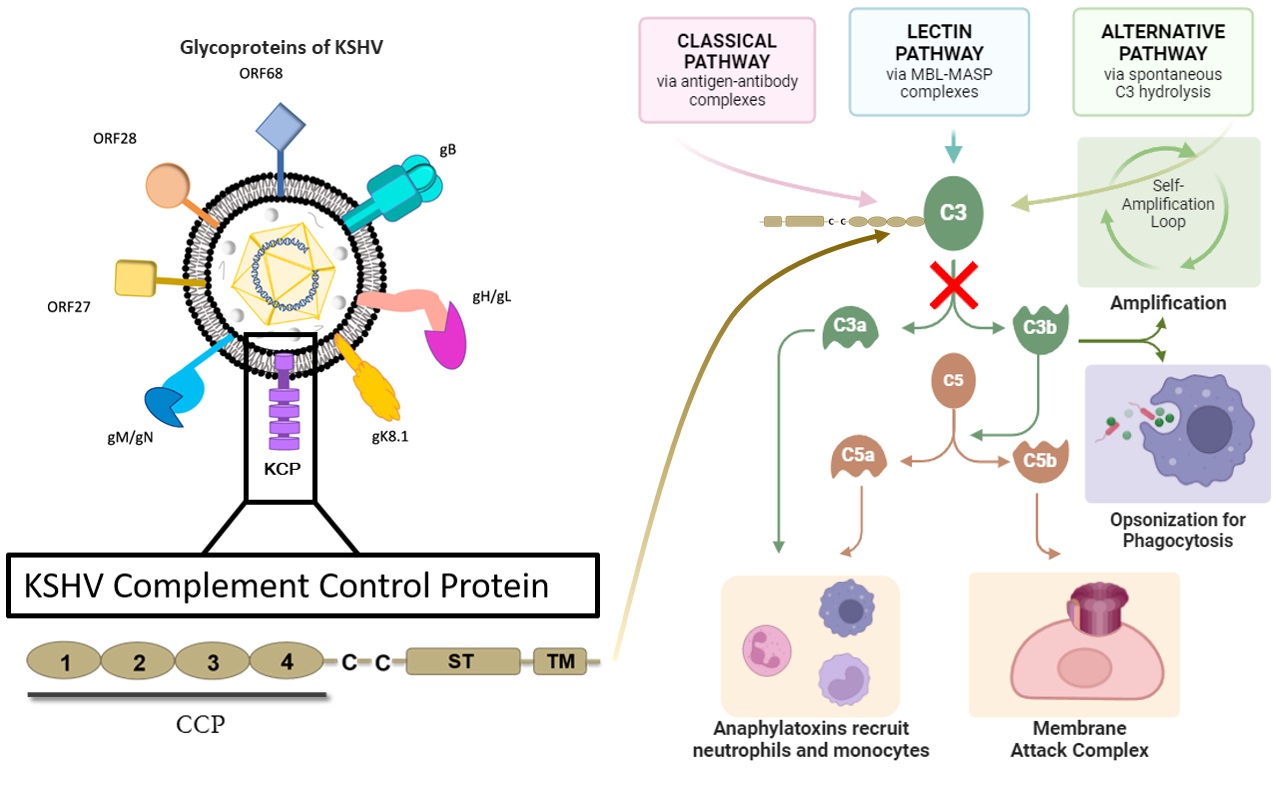
A08 – Structural Characterization of Complement Control Protein of Kaposi´s Sarcoma Associated Herpes Virus
Hera Fatima
associated
Universität zu Lübeck (UzL)
–
–
–
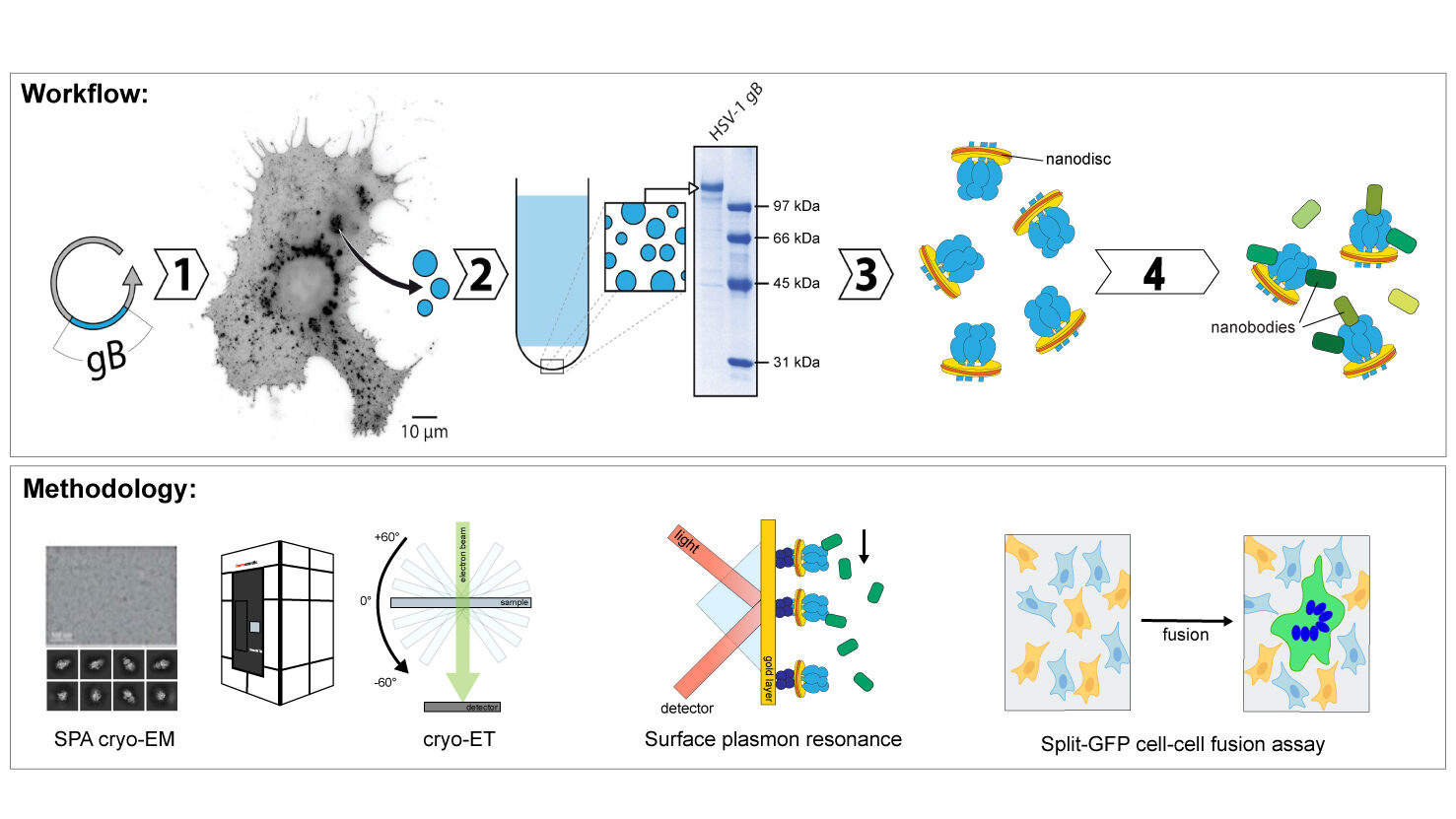
B01 – High-resolution structure and conformational flexibility of Herpes simplex virus 1 (HSV-1) fusion protein gB
Julia Nentwig
Centre for Structural Systems Biology (CSSB)

B02 – Structural and functional characterisation of the scavenger receptor B1 – a cellular receptor for Hepatitis C virus (HCV)
Niklas Ebersberger
Universität zu Lübeck (UzL)

B03 – Topological and functional characterization of the pestiviral non-structural protein 4B (NS4B)
Sabine Höppner
associated
Universität zu Lübeck (UzL)
–
–
–

C01 – Structural and functional features of a conserved DNA-binding domain in herpes viral proteins as a basis for innovative targeted therapies
Fatama Sornaly
Universität zu Lübeck (UzL)

C02 – Characterisation of the structural and molecular mechanism how Merkel cell polyomavirus (MCPyV) Large T-Antigen tumps hallmark mutations occur
Tommaso Mari
University Medical Center Hamburg-Eppendorf (UKE)

C03 – Structural and functional characterisation of the hepaci- and pestiviral NS2-NS3 polyprotein region
tba
Universität zu Lübeck (UzL)
–
–
–

C04 – Role of the human cytomegalovirus (HCMV) E3 SUMO ligase pUL69 in virus reactivation
Sairam Dasika
Universität zu Lübeck (UzL)
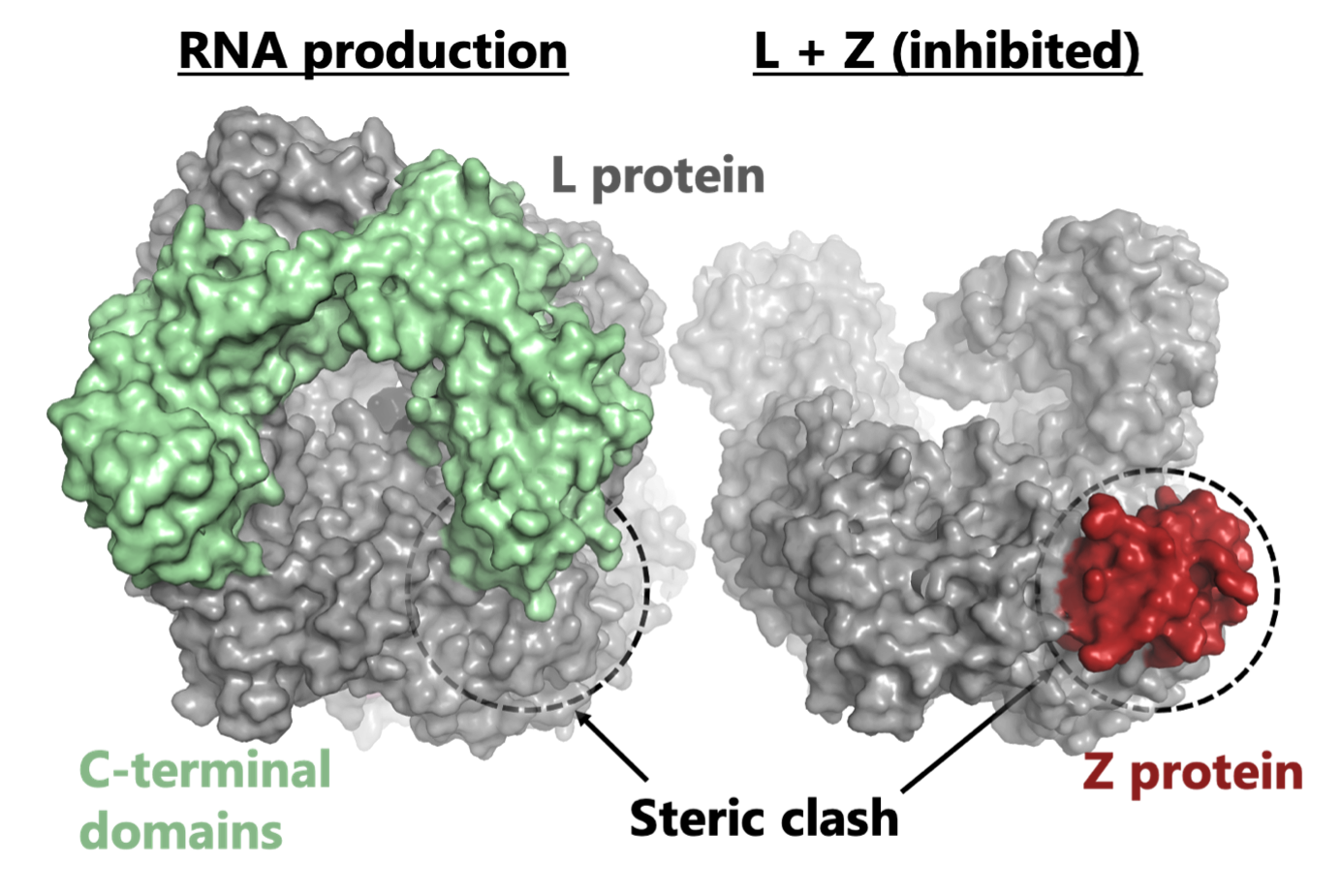
C05 – The mechanism of Z-based RNA synthesis inhibition in Lassa virus as an antiviral strategy
Annika Rammelt
Centre for Structural Systems Biology (CSSB)
–
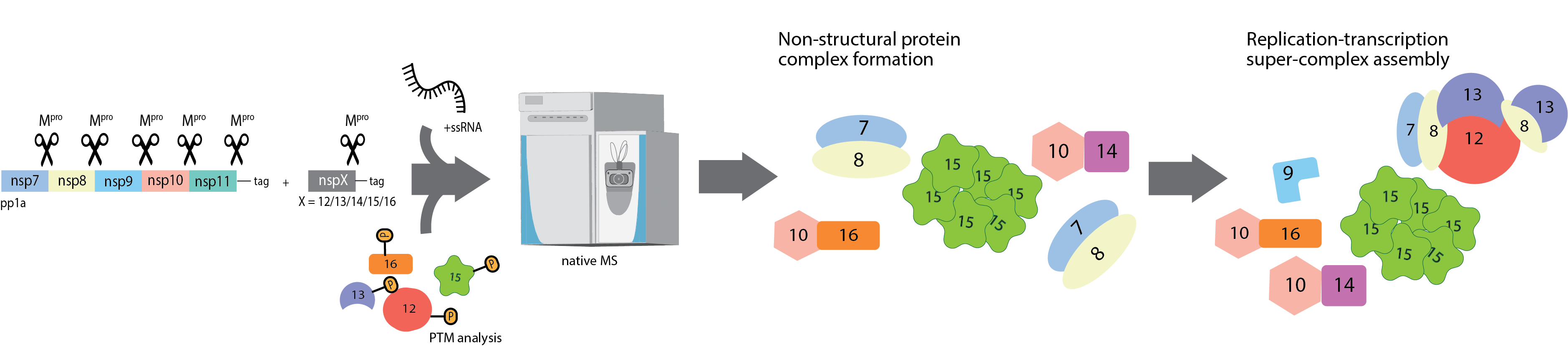
C06 – Revealing SARS-CoV-2 replication-transcription super-complex (RTC) architecture and dynamics via native MS
Fatema-Aqila Said
associated
Centre for Structural Systems Biology (CSSB)
VISION Research Projects – 2nd cohort

A01.2 – Targeting viral Achilles heels with AI-driven protein design
Aria Dang
Centre for Structural Systems Biology (CSSB)

A02.2 –
Stefanie Rössler
Centre for Structural Systems Biology (CSSB)

A03.2 –
tba
Universität zu Lübeck (UzL)
–
–
–

A04.2 – Modeling interfaces and designing peptides and other molecules to disrupt capsid assembly, stability, and size
Shaina To
Centre for Structural Systems Biology (CSSB)
–
–
–

B01.2 – Comprehensive structural investigation of the herpesvirus viral membrane glycoproteins
Iker Arriaga Saez
Centre for Structural Systems Biology (CSSB)

B02.2 – Structural and Functional Characterization of Intact Scavenger Receptor Class B Type 1
Alexander Müller
Universität zu Lübeck (UzL)

B03.2 – Interplay between Hepatitis C Virus and the hepatic lipid droplet mosaic
Muhammad Nadeem
associated
Universität zu Lübeck (UzL)
–
–
–

C01.2 – Structural investigation of UL3 from herpes simplex virus type 1: A structural homolog of a conserved DNA-binding protein domain in herpesviral proteins?
Anna Weissenburg
Universität zu Lübeck (UzL)

C02.2 – Exploring the interactions and tethering between Merkel cell polyomavirus (MCPyV) genome and host cellular chromatin
Vishakha Ramamurthy
University Medical Center Hamburg-Eppendorf (UKE)

C03.2 – Structural and functional characterization of the pestiviral NS2pro-NS3pro polyprotein region
Malte Morische
Universität zu Lübeck (UzL)
–
–
–

C04.2 – The role of the non-classical deSUMOylase LUNA from HCMV in viral reactivation from latency
Jasper Kroes
Universität zu Lübeck (UzL)

C05.2 – tba
tba
Centre for Structural Systems Biology (CSSB)

C06.2 – Dissection of the pestiviral replicase and packaging complexes by proximity labelling and biorthogonal crosslinking approaches
Yasmin Roeder
associated
Universität zu Lübeck (UzL)
–
–
–

C07.2 – Discovery and characterization of small-molecule inhibitors targeting the L protein of arena- and phenuiviruses
Simon Rothbauer
associated
Centre for Structural Systems Biology (CSSB)

C08.2 – Towards assembly and remodelling of Lassa virus ribonucleoparticles in vitro
Morten Schumann
associated
Centre for Structural Systems Biology (CSSB)
